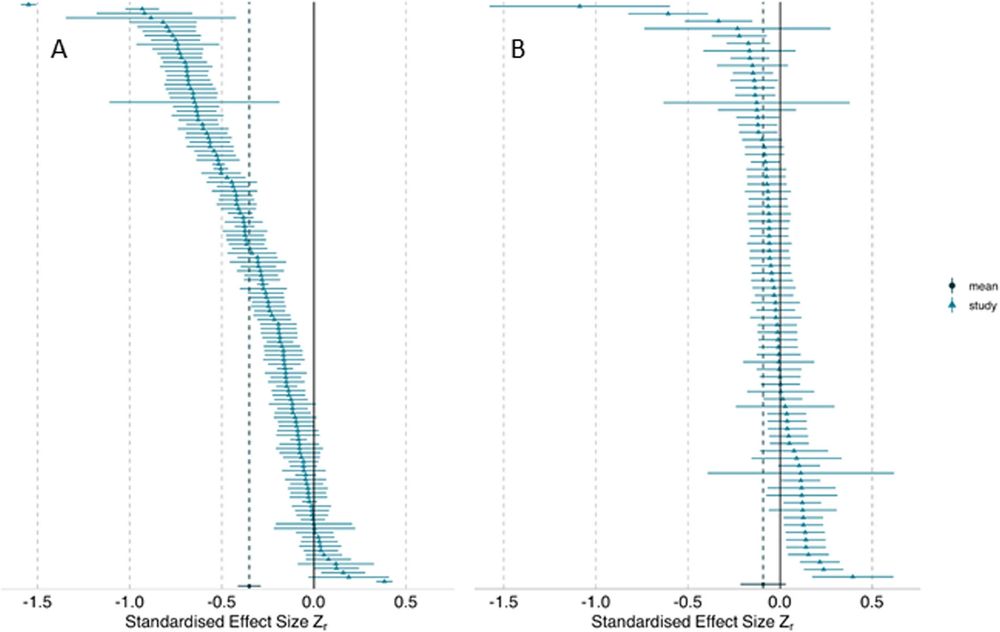
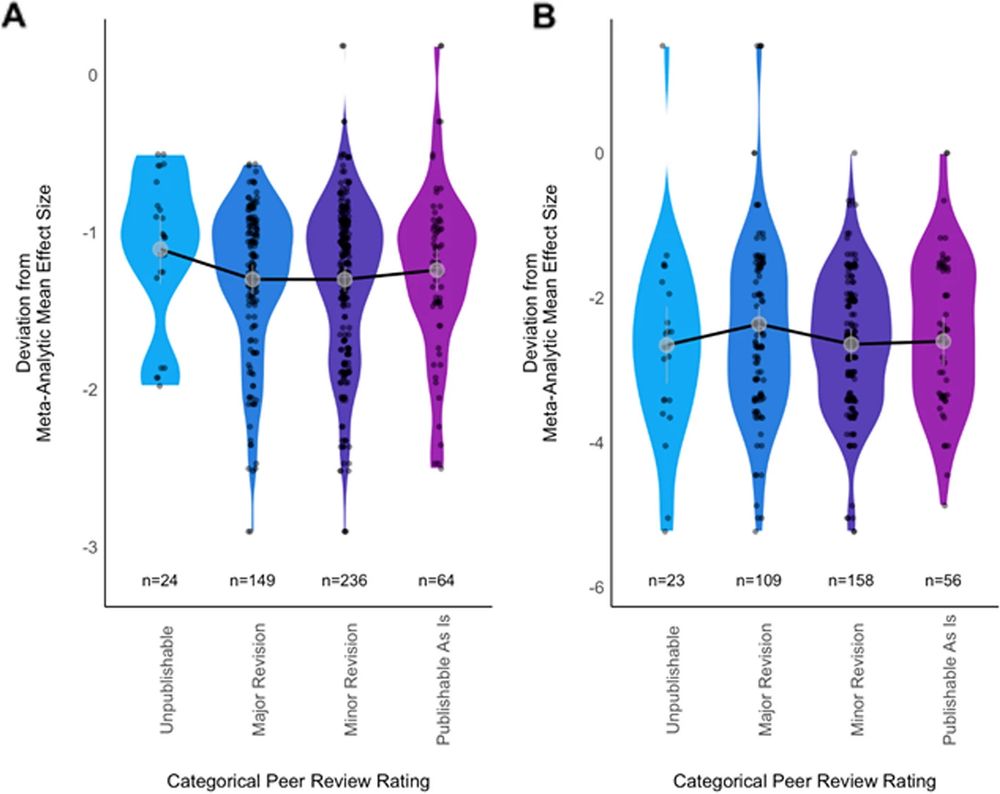
This Registered Report masterpiece just dropped at BMC Biology, brilliantly led by a great team with the help of 300+ analysts & reviewers
Same question, same data: go figure!
tl;dr: Substantial heterogeneity among results comes from differences among analytical choices
🔗 doi.org/10.1186/s129...
07.02.2025 04:00
👍 104
🔁 52
💬 2
📌 3

The first many-analysts study in ecology is finally published! 🥳🙌
300+ coauthors and 5+ years, this was a massive effort by @elliotgould.bsky.social Hannah Fraser Tim Parker and co.
Open access 👉 bmcbiol.biomedcentral.com/articles/10....
06.02.2025 17:57
👍 153
🔁 64
💬 2
📌 7
Thanks to the GitHub community discussion I've managed to solve the problem! Yay for the power of open-source code dev. Check out the package-down page for the ManyEcoEvo R package. Needs some articles / vignettes added, but it's looking good :D egouldo.github.io/ManyEcoEvo/i...
07.11.2023 08:57
👍 2
🔁 0
💬 0
📌 0

Marcia Langton: ‘Whatever the outcome, reconciliation is dead’
The referendum campaign has cemented racism into the body politic, and the ‘baseless’ rejection of ‘Yes’ will create a bleak future for Australia and those who stood with First Nations people.
"Both major parties say they support the recognition of Indigenous Australians. This is not true in practice. In fact, the appearance of policy agreement on Indigenous constitutional recognition is a saga of deceit and treachery, kicked down the road for more than a decade."
13.10.2023 23:13
👍 7
🔁 3
💬 1
📌 0
This by Sarah Schiavone, @simine.com and @kimberlyquinn.bsky.social might be what you were thinking of: osf.io/preprints/ps.... I highly recommend seaboat.io, it's been a useful reviewer tool (and a nice complement to the blogpost of mine that @matthewwinn.bsky.social linked (thank you Matthew!).
14.10.2023 20:51
👍 5
🔁 1
💬 1
📌 1
As Brian put, the contingency of results and conclusions on analytic heterogeneity is rarely acknowledged and is under-appreciated in practice. Our paper seeks to bring this issue to the attention of ecologists and evolutionary biologists. 2/
15.10.2023 05:21
👍 4
🔁 0
💬 0
📌 0
I don't believe science is broken, and that's not argued in the paper. We should always be striving to improve scientific practice, and our paper is a part of that task. We did not expect a single result or conclusion, which is why we preregistered models examining potential causes of variation. /1
15.10.2023 05:20
👍 2
🔁 0
💬 0
📌 0
Preregistering multiple models could help contextualise inference of a preferred model accounting for analytic uncertainty, "If we had chosen an alternative specification, how might this influence our results and conclusions"? /2
15.10.2023 05:01
👍 1
🔁 0
💬 1
📌 0
Multiverse analyses/many-analyst analyses/specification curve analyses highlight the many researcher degrees of freedom present within a typical study, as we demonstrated. In this context, preregistration can mitigate undisclosed exploitation of these RDF's. 1/
15.10.2023 05:00
👍 3
🔁 0
💬 1
📌 0
Here's the link to the pre-print doi.org/10.32942/X2G..., the link above is to the Nature news article covering the paper.
15.10.2023 04:50
👍 2
🔁 0
💬 0
📌 0
That's from our preprint doi.org/10.32942/X2G.... And correct, our argument here is about preregistration of multiple models as an alternative to analysing a single model. And absolutely, only biologically meaningful models should be preregistered.
15.10.2023 04:49
👍 2
🔁 0
💬 0
📌 0
Our intended point here was that *multiple* models, rather than a single model or analysis, could be preregistered. Both preregistration and robustness analyses are offered as alternatives to 'single analyses', as is stated in the topic-sentence shown in the screenshot.
15.10.2023 04:29
👍 5
🔁 0
💬 0
📌 0
Many EcoEvo Analysts has been years in development with HUNDREDS of contributors from ecology analysing (and reviewing analysis) the same two datasets. Herioc effort by MetaMelb's Elliot Gould & Hannah Fraser + Tim Parker, Shinichi Nakagawa, Pete Vesk and many others ecoevorxiv.org/repository/v...
05.10.2023 03:25
👍 15
🔁 13
💬 1
📌 0




