

Attending the Scipion workshop at KAIST. Looking forward to learning a lot over the next 4 days


Attending the Scipion workshop at KAIST. Looking forward to learning a lot over the next 4 days
Congrats!!!

Talented researchers from Rho lab at Seoul National University just published impressive results.
Glad I could help with some of the cryo-EM data collection for this work.
www.science.org/doi/10.1126/...
Congratulations, Oli!
Truly fantastic work. The more small protein research, the better!
Also, please say hello to Kookju as well.
I carried out over the past two years. It’s very disappointing that my contribution leading to this achievement wasn’t recognized.
That said, I wish the authors and everyone involved the best of luck. If you’re interested in the paper, please reach out to the corresponding authors listed there.
I think this will be my last post on this topic. I don’t want to keep talking about the authorship issue. Even though I wasn’t directly involved in the experiments/analyses for the paper, I believe this achievement wouldn’t have been possible without the discussions I led on SNS and the work
The structures and maps in the paper have been newly deposited.
I was also told that the author list consists of people who directly participated in the actual paper writing, and they believe this is the right approach.
I contacted the first author and received specific reasons:
The data was newly processed after I left the company, and there is additional data as well, with ongoing data collection planned for better results.
At this point, it seems like including my name in the author list wouldn’t have much meaning. I just want to hear from the paper authors why I had to be excluded from this result. 3/3

After receiving this response, I'm so bewildered that I can't think straight. Looking at the paper's author contributions, 'H.K.' is included. Of course, there's no name corresponding to H.K. in the author list. 2/3
Just received this response about why I was excluded from authorship on the small protein cryo-EM paper "We have to respect the opinions of the members who actually conducted the research. What they confirmed is that the research was conducted after Dr. Kim left the company." 1/3

PNCC Cryo-EM modeling and validation workshop on Sep 3-5, 2025. Register now! tinyurl.com/Cryo-EMModel...
We tried to run RELION but encountered problems during data processing.
I’m not sure if some of the authors have much experience with RELION, but I can ask about that.
As you said, PLK1 is probably too small, making particle alignment very difficult.
Anyway, I will discuss this with the authors about how to address the issue. I will keep you updated.
Once again, thank you.
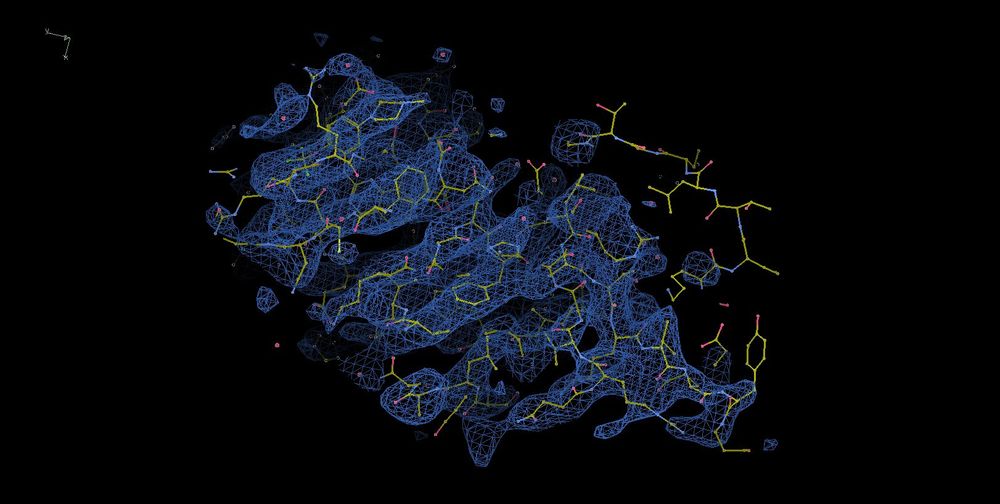
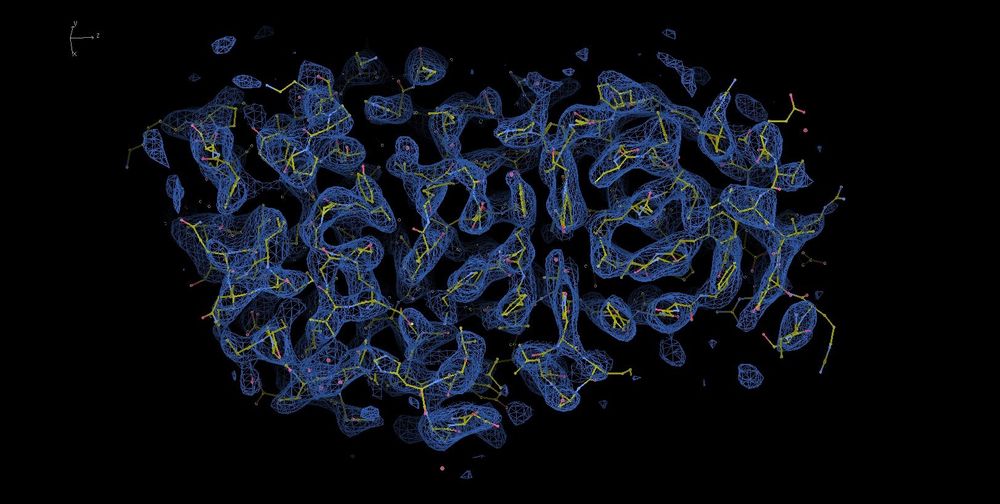

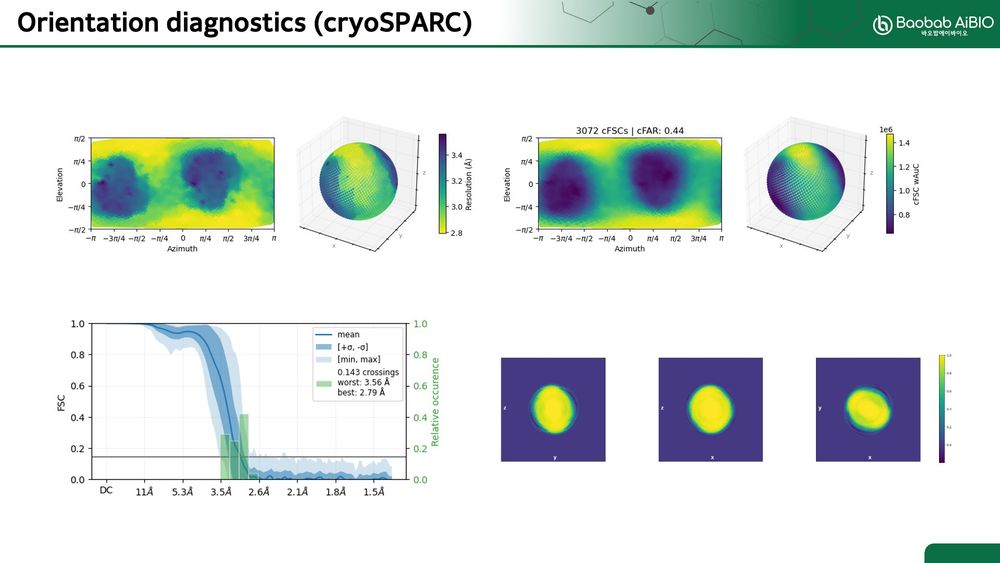
Thanks for the kind words, Oli.
The last time I processed the PLK1 map, it looked like the images I attached.
We also added 30° tilted micrographs for processing. It improved the orientation diagnostics results. However, the map is still anisotropic.

but I could not finish any of them and it looks like they wrote the paper with the same data I was working on.
Anyway, I will contact the authors to bring up some issues discussed here.
BTW, MBP structure is available.
www.rcsb.org/structure/8YBE
@olibclarke.bsky.social
@jhschaef.bsky.social
When I was at BaobabAiBIO, I was hesitant to publish the results because the PLK1 map was incomplete (I wrote about my concerns on Twitter).
Before quitting my job, I tried to finish at least these 2 things:
- getting a complete PLK1 map
- uploading MBP/PLK1 data to EMPIAR
BTW, Jan-Hannes Schäfer @jhschaef.bsky.social and Gabriel C. Lander @landerlab.bsky.social provided an excellent PREreview of the paper. I really appreciate it.
prereview.org/reviews/1581...

Although I’m not on the author list, the MBP/PLK1 cryo-EM structure paper has been published on bioRxiv.
FYI, I was not involved in any writing or reviewing of the paper
www.biorxiv.org/content/10.1...

I’m experiencing repeated errors during tomography data acquisition.
Any suggestions on how to resolve this issue? Thanks in advance.

Thrilled to see my former lab's paper on bioRxiv. Their graphene-based affinity grid successfully resolved low-abundance protein complexes. After such a long journey, it's wonderful to see this project finally producing good results!
www.biorxiv.org/content/10.1...
As we head into 2025, let's take one final look back at 2024 and how the year unfolded in the PDB archive. There were 15319 entries released last year but what more can we find out about those structures ⚖️?
A thread 1/11 🧵




Snow in Seoul

Come work with us! Only a few days remaining to apply for the #cryoET / #CLEM facility scientist position at the @icrlondon.bsky.social! #cryoEM
jobs.icr.ac.uk/vacancies/96...
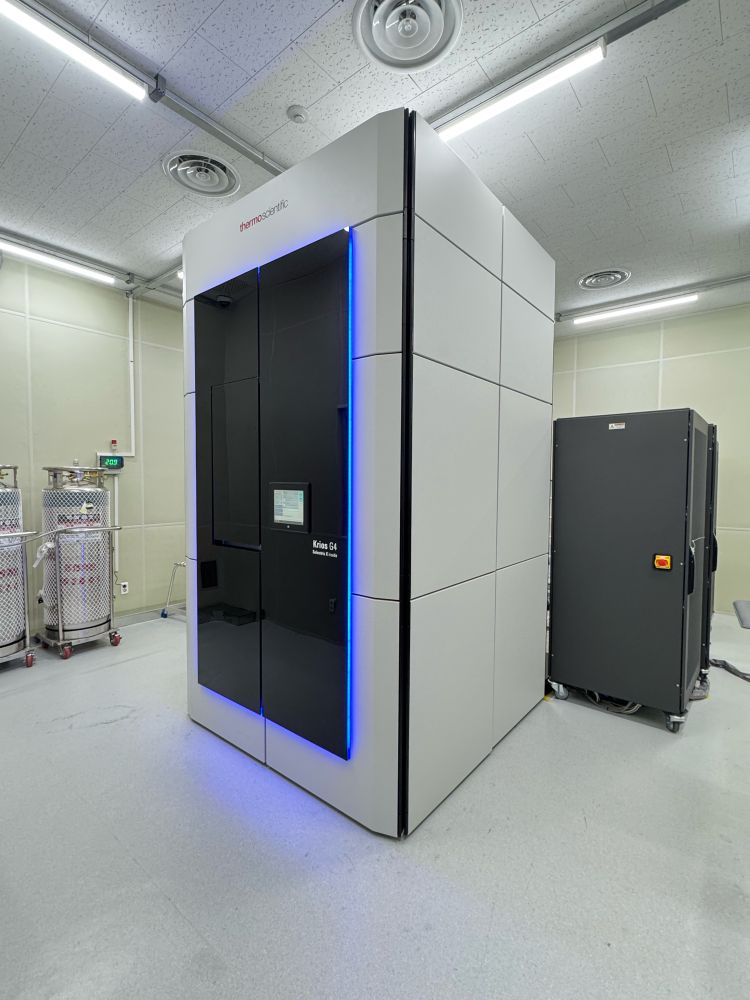


New office desk & Krios G4 at SNU NCIRF
Starting from September, I will begin working at the Core Facility for Cell and Macromolecule Imaging and the National Center for Inter-university Research Facilities at Seoul National University to manage the cryo-EM equipment and provide research support.

Update on our kinase domain side project (37 kDa with a ligand): Previous sample had severe preferred orientation issue, pausing the project. Now back at it with new 2D classification results. Still preferred orientation persists, but we're optimistic about getting a map. keep the fingers crossed.
Our MBP cryo-EM structure is released.
Next uploading the micrographs to EMPIAR.
www.rcsb.org/structure/8YBE

Join us for the 2024 Cryo-EM Workshop for the pharma and biotech industries! Dive into the latest trends, techniques, and applications of Cryo-EM with experts from Baobab AiBIO and Thermo Fisher Scientific. (conducted in Korean / industry only) Mar 14, 10:00-17:00. docs.google.com/forms/d/e/1F...
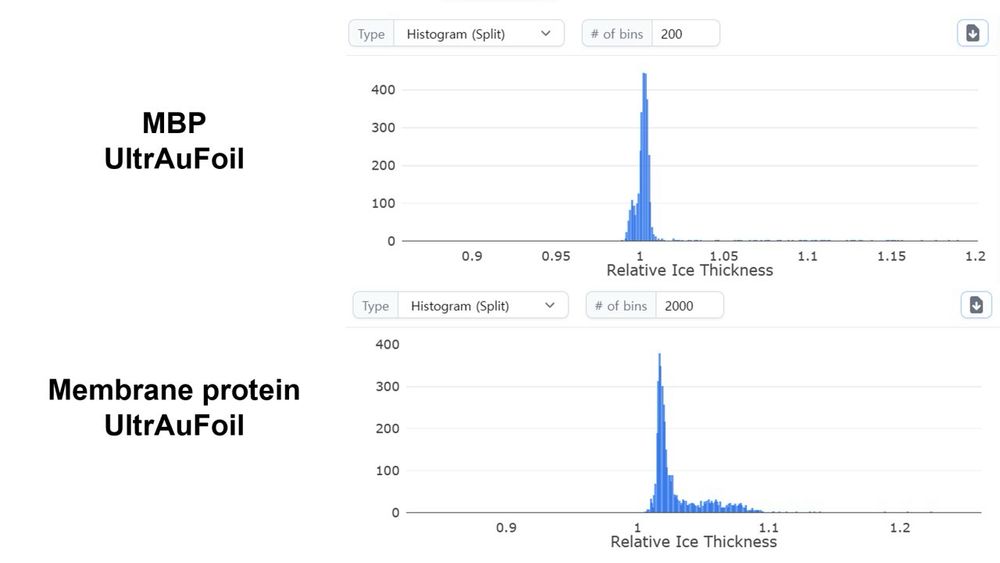
Oli B. Clarke (I’m mot sure he is using bluesky or not) asked about the ice thickness of our MBP dataset. So I checked the relative ice thickness plot in cryoSPARC and compared our membrane protein dataset. Here is the result.