How does the piRNA pathway solve the self vs. non-self problem? 🧬
Since piRNAs come from single-stranded RNA, how does the cell choose the right ones? For years, "piRNA clusters" were seen as THE privileged source. But are they really special and earmarked for biogenesis? (1/19)
13.02.2026 15:11
👍 90
🔁 51
💬 2
📌 4
A Naïve RNA Sampling Core Enables Adaptive piRNA Specificity Against Transposable Elements https://www.biorxiv.org/content/10.64898/2026.02.07.704324v1
10.02.2026 05:17
👍 15
🔁 11
💬 0
📌 2

Ulrich’s research will help advance our understanding of RNA biology & gene regulation, as well as how these mechanisms might be altered during viral infections & other diseases.
@hohmannulrich.bsky.social comes from the @impvienna.bsky.social / @imbavienna.bsky.social
www.imb.de/about-imb/ne...
09.02.2026 11:14
👍 40
🔁 16
💬 1
📌 1

Programmatic design and editing of cis-regulatory elements
The development of modern genome editing and DNA synthesis has enabled researchers to edit DNA sequences with high precision but has left unsolved the problem of designing these edits. We introduce Le...
After a huge amount of work w/ @alex-stark.bsky.social's group, a new version of our Ledidi preprint is now out!
In an era of AI-designed proteins, the next leap will be controlling when, where, and how much of these proteins are expressed in living cells.
www.biorxiv.org/content/10.1...
10.12.2025 15:18
👍 60
🔁 26
💬 2
📌 0
Happy to share that my PhD project is finally published!🪱✨
Selfish genes are found across the tree of life. They can disrupt inheritance patterns and at the same time act as units for molecular innovation. Here we tried to answer one big question: how do selfish genes emerge in the first place?
24.11.2025 21:10
👍 75
🔁 36
💬 3
📌 0
🪱 Selfish genes are everywhere and drive some of biology’s biggest innovations (CRISPR, antibody recombination, epigenetics). Yet almost no one asks the obvious question: how does a selfish gene begin? Our new manuscript uncovers how selfishness can emerge directly from the host genome.
24.11.2025 13:03
👍 61
🔁 38
💬 1
📌 1
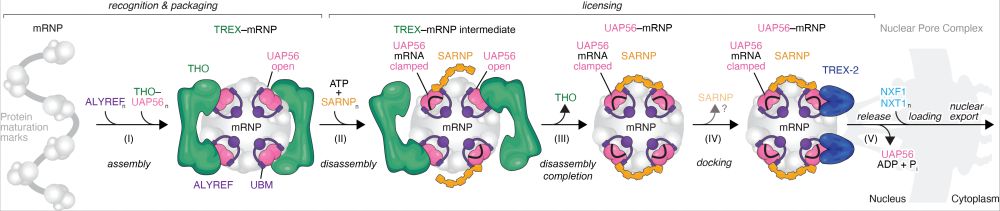
Finally out in @nature.com! We uncovered a mechanistic framework for a general and conserved mRNA nuclear export pathway. www.nature.com/articles/s41.... 1/
19.11.2025 23:21
👍 106
🔁 42
💬 3
📌 1
Research
IMB Mainz
Lastly, I’m excited to join @imbmainz.bsky.social in 2026 to start my own lab. We'll explore new mechanisms in eukaryotic gene expression, leveraging ‘evolutionary play’ to uncover how regulation, repurposing, and hijacking shape RNA biology. tinyurl.com/y4x29ctt
Thanks for reading! 20/20
19.11.2025 23:21
👍 19
🔁 5
💬 0
📌 1
just in time for the opening of the @hohmannulrich.bsky.social group at @imbmainz.bsky.social
what started as a project on how cells export piRNA precursors, ended up as a tour de force in mRNA export. truly wonderful collaboration with @plaschkalab.bsky.social at the @viennabiocenter.bsky.social
07.11.2025 18:03
👍 32
🔁 11
💬 0
📌 0

Graphical abstract: The Drosophila OSC Genome as a resource for transposon and piRNA biology. The figure illustrates the workflow and key findings. Left: De novo genome assembly using Oxford Nanopore Technologies (ONT) long reads and Hi-C data generates a phased assembly distinguishing unique (blue) and repetitive (orange) sequences. Dot plot comparison between OSC-r1.01 and dm6 reference genomes shows overall synteny with extensive structural variation. Middle: A freely accessible UCSC genome browser session displays multi-omics data tracks including gene models, transposon insertions, chromatin accessibility, transcription, small RNAs, and histone modifications. Right: New insights into flamenco piRNA cluster biology reveal >730 kb transcribed from a single promoter without major splicing. Tethering assays demonstrate long-range silencing effects across the locus, and genome browser tracks show coordinated regulation of piRNA production, transcription, and chromatin state. This resource enables comprehensive studies of transposon regulation and piRNA pathway function in a widely-used Drosophila cell line.
When transposons jump, genomes diverge - even in cultured cells.
I am happy to share our new preprint: a chromosome-scale genome assembly for Drosophila OSC cells, one of the key model systems in the piRNA field, especially for nuclear piRNA biology. 🧬🧵 (1/12)
14.10.2025 20:34
👍 66
🔁 24
💬 3
📌 4
My first first-author paper is out!🎉
Here we propose a model where a silencing complex, PIWI*, assembles on target RNAs to recruit effectors and shut down transposon activity.
Huge thanks to the Brennecke and Plaschka labs, especially Julius and Clemens, and all co-authors!
17.09.2025 13:00
👍 48
🔁 20
💬 3
📌 1
12/ This work is part of my PhD project at the @viennabiocenter.bsky.social. A big Thank You to @juliusbrennecke.bsky.social and Kirsten Senti for their supervision! To my co-authors Liudmila and @86dominik.bsky.social, and to the rest of the Brennecke lab!
14.08.2025 17:09
👍 5
🔁 0
💬 0
📌 0
11/ All in all, our work strongly suggests that the decision to process a transcript into piRNAs is not a feature of the genomic source locus but rather a decision that is taken in the cytoplasm after export of a TE antisense-containing transcript.
14.08.2025 17:09
👍 5
🔁 0
💬 1
📌 0

Evolution of KoRV-A transcriptional silencing in wild koalas
Koala retrovirus-A (KoRV-A) is spreading through wild koalas in a north-to-south wave while transducing the germ line, modifying the inherited genome …
9/ Importantly, this is not restricted to flies, Bill Theurkauf and colleagues, showed that silencing of the invading KoRV-A retrovirus in wild koala populations strongly correlates with the presence of a single antisense insertion in a host gene 3′ UTR. www.sciencedirect.com/science/arti...
14.08.2025 17:09
👍 4
🔁 1
💬 1
📌 0
8/ Altogether, our data show that antisense insertions of TEs in host gene exons are sufficient to elicit a piRNA response, which expands the repertoire of piRNA source loci to host genes expressed in gonads in addition to “classical” piRNA clusters.
14.08.2025 17:09
👍 3
🔁 0
💬 1
📌 0

A GFP transgene containing an antisense tirant insertion in the 3'UTR can silence tirant expression. An insertion in the sense orientation does not affect the expression of tirant.
7/ To test this idea, we generated a synthetic construct in which we placed antisense tirant sequence into the 3′ UTR of a UAS-GFP transgene. When expressed in follicle cells, this transgene silenced tirant expression, confirming our model.
14.08.2025 17:09
👍 2
🔁 0
💬 1
📌 0
6/ This led us to propose a model in which antisense piRNAs can only be generated if a TE inserts itself into a host transcription unit (gene or piRNA cluster). This generates a cytoplasmic antisense RNA that is processed into piRNAs.
14.08.2025 17:09
👍 5
🔁 0
💬 1
📌 0

An insertion of tirant in the antisense orientation in the 3'UTR of the gene Fs(2)Ket leads to the production of piRNAs.

An ectopic copy of the gene Fs(2)Ket containing the tirant insertion is sufficient for the production of piRNAs and silencing of tirant. This requires the production of a chimeric transcript containing both the Fs(2)Ket and the tirant RNA.
5/ Surprisingly, we found that a single antisense insertion of tirant in the 3′ UTR of a host gene, Fs(2)Ket, is sufficient to trigger piRNA production and silence tirant. piRNA production from this insertion requires host gene transcription and a chimeric mRNA-TE transcript.
14.08.2025 17:09
👍 3
🔁 0
💬 1
📌 1
4/ Some strains lack tirant insertions in flamenco or in any other known piRNAs cluster. However, they are still able to produce piRNAs and silence tirant. So where are these piRNAs coming from?
14.08.2025 17:09
👍 4
🔁 0
💬 1
📌 0

The tirant-GFP:lacZ reporter is expressed in the somatic follicle cells of the Drosophila melanogaster ovary and is silenced by piRNAs.
3/ We first show that tirant is exclusively expressed in somatic cells of the ovary. Here, most strains also produce tirant piRNAs from tirant insertions in flamenco, the major piRNAs cluster active in that tissue. But this is not true in all strains …
14.08.2025 17:09
👍 4
🔁 0
💬 1
📌 0
2/ Over the past century, several TEs invaded Drosophila melanogaster (thanks to the work of @rokofler.bsky.social). We traced how natural strains acquired piRNA immunity against one of these TEs, tirant, an LTR retrotransposon.
14.08.2025 17:09
👍 5
🔁 0
💬 1
📌 0
1/ How do animals develop immunity against a newly encountered transposable element from scratch? Our study reveals that the mobility of TEs is their Achilles heel, allowing hosts to develop a powerful small RNA-mediated silencing response.
www.biorxiv.org/content/10.1...
14.08.2025 17:09
👍 49
🔁 27
💬 4
📌 3










